Cells can be understood as highly coordinated systems in which DNA functions as a comprehensive but inert blueprint, requiring precise interpretation to become biologically active. Gene expression depends on regulatory proteins that orchestrate transcription across time and space. Within this framework, the Bromodomain and Extra-Terminal (BET) family plays a pivotal role. While BRD4 has traditionally been emphasized for its function in transcriptional elongation, emerging research highlights a distinct and essential contribution of BRD2. In mouse embryonic stem cells, BRD2 stabilizes RNA polymerase II at gene promoters via interaction with TFIID, particularly under conditions where pause release is compromised. Additionally, MOF-mediated histone H4 acetylation enhances BRD2 recruitment and chromatin organization.
BRD2: The Emergency Brake That Prevents Transcriptional Collapse
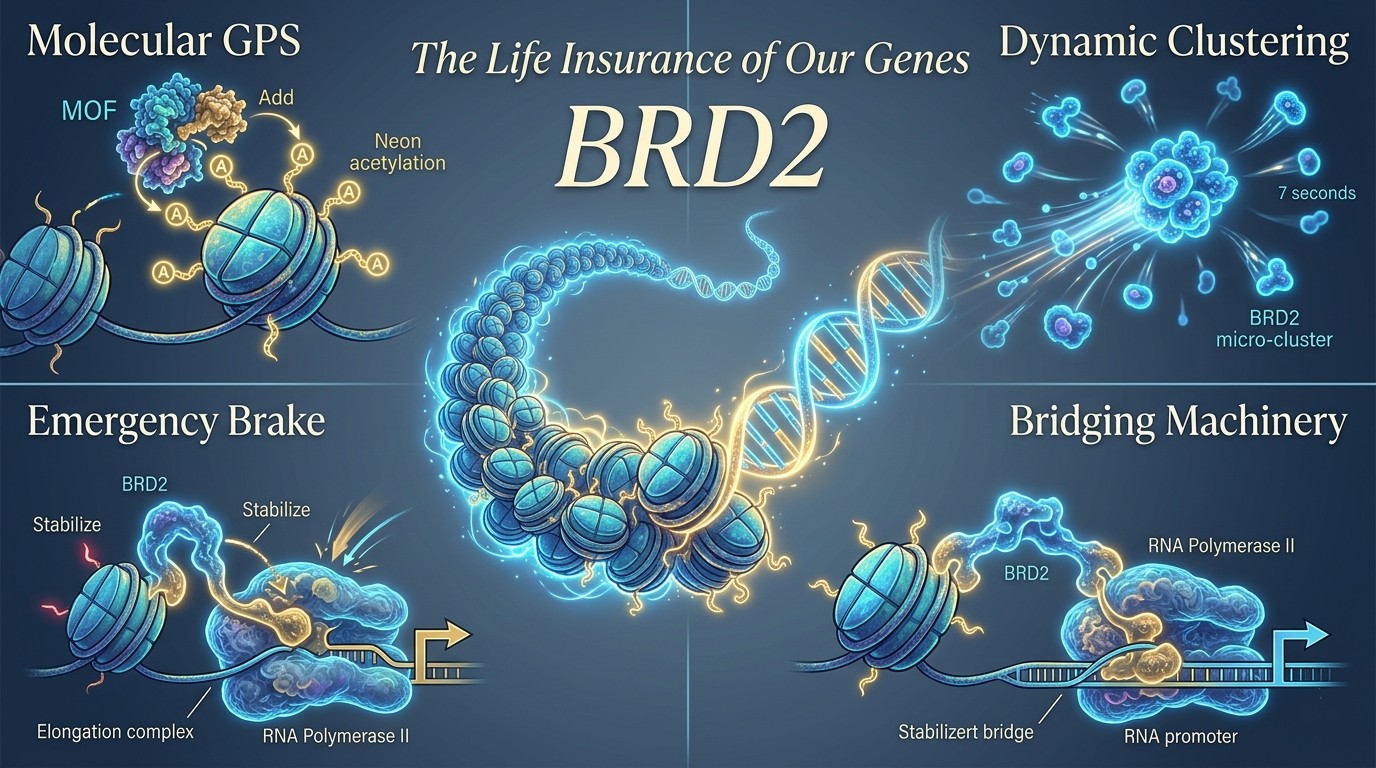
For many years, the loss of BRD4 was considered sufficient to severely disrupt cellular transcription by impairing RNA synthesis. However, recent studies (Erdogdu et al., 2026) reveal that BRD2 acts as a molecular safeguard, becoming particularly important when transcriptional elongation is compromised. This role is especially evident under pharmacological inhibition with compounds such as flavopiridol, which block transcriptional pause release. Under these conditions, BRD2 is increasingly recruited to gene promoters, where it helps maintain occupancy of RNA polymerase II. Whereas BRD4 primarily promotes productive elongation, BRD2 stabilizes the initiation stage, preventing premature dissociation of the transcriptional machinery and preserving transcriptional competence during cellular stress.
The Molecular “GPS”: MOF-Mediated Histone Acetylation
How does BRD2 identify its precise landing sites across the vast genomic landscape? The answer lies in histone acetylation, a chemical modification that functions as a molecular positioning system. The lysine acetyltransferase MOF operates as a high-precision navigator, depositing acetyl marks on histone H4 that define transcriptionally active regions. Among these marks, H4K16ac plays a dominant role, although MOF also contributes to H4K5ac, H4K8ac, and H4K12ac deposition. Loss of MOF activity produces a dramatic disruption in chromatin organization, reducing BRD2 occupancy across thousands of genomic loci, over 9,500 binding sites, while exerting comparatively minor effects on related BET proteins such as BRD3 and BRD4. This striking specificity highlights a dedicated functional partnership between MOF and BRD2 during transcriptional initiation. Together, their acetylation-dependent spatial dynamics coordinate RNA polymerase II recruitment and stabilize the early stages of gene activation.
Dynamic Clustering: Seven Seconds of Molecular Agility
The spatial organization of transcription factors within the nucleus reveals a striking level of dynamic engineering. Using time-correlated photoactivation localization microscopy (tcPALM), researchers can now visualize molecular interactions that were previously beyond optical resolution. Unlike BRD4, which forms relatively stable transcriptional condensates, BRD2 operates through highly transient micro-clusters that assemble and dissolve within seconds. These clusters display lifetimes of approximately 6.5–7.5 seconds, reflecting the flexible nature of BRD2’s intrinsically disordered region (IDR). Rather than representing disorder, this structural flexibility enables rapid molecular responsiveness, allowing BRD2 to reorganize dynamically and adapt transcriptional initiation processes to changing cellular demands.
Bridging the Core Machinery Connecting to the Transcriptional Engine
The final act of BRD2 function is both physical and mechanical: it serves as a molecular bridge linking chromatin signals to the core transcriptional machinery. Central to this role is its interaction with TFIID, a large multiprotein complex responsible for recognizing promoter sequences and establishing transcription start sites. In particular, BRD2 associates with the TAF1 subunit, reinforcing the assembly and stability of the pre-initiation complex. Rather than simply recruiting RNA polymerase II, BRD2 enhances its retention at promoters, ensuring sustained occupancy during early transcriptional stages. This distinction is critical; successful transcription requires not only arrival but persistence. By anchoring TFIID components and supporting RNA polymerase II positioning, BRD2 stabilizes the transcriptional platform, enabling cells to maintain gene activation potential even under fluctuating or stress-induced transcriptional conditions.
Toward a New Era of Precision Epigenetic Medicine
Distinguishing the initiation-focused role of BRD2 from the elongation-driven function of BRD4 reshapes our understanding of transcriptional control and opens new therapeutic possibilities for diseases such as cancer and inflammatory disorders. Historically, BET inhibitors targeted the entire protein family broadly, often disrupting multiple regulatory layers simultaneously. These new insights suggest a shift toward precision epigenetic strategies, where drugs could selectively modulate BRD2-dependent clustering or recruitment without compromising global transcriptional balance. At any given moment, within every living cell, transient BRD2 clusters assemble and dissolve in rapid cycles, sustaining promoter readiness and preserving transcriptional responsiveness. The future of molecular medicine may therefore extend beyond correcting genetic sequences, focusing instead on fine-tuning the dynamic regulators that orchestrate gene expression with remarkable temporal and spatial precision.




Discussion